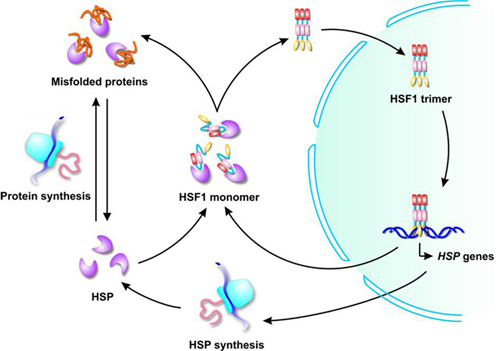
This review summarizes fundamental understanding and recent advances in multidomain protein folding and mechanistic insights into molecular chaperones. In this review, we focus on multidomain protein folding, assisted by molecular chaperones, and discuss how molecular chaperones alter the folding properties of client proteins. This activity switching of molecular chaperones is believed to be important in regulating the folding of multidomain proteins. For example, one chaperone can work as a foldase for one group and as a holdase for another group of client proteins. One chaperone can exert multiple different activities, depending on the conditions or client proteins. Interestingly, chaperone activity is rather variable. 
Furthermore, some molecular chaperones, including caseinolytic mitochondrial matrix peptidase chaperone (Clp)/heat shock protein 100 (Hsp100) chaperones, are classified as “Unfoldases” that unwind the structure of damaged proteins. The holdase activity of molecular chaperones is known to be important for protein folding, in which the chaperone slows down the folding process for higher yield. For instance, trigger factor (TF), DnaK, and Spy work as foldases to promote client protein folding SecB and heat shock protein 40 (Hsp40) work as holdases to inhibit client protein folding. “Holdases” prevent or retard client protein folding, often for efficient protein translocation. “Foldases” assist in the folding of newly synthesized proteins. 
Molecular chaperones can be classified based on their characteristic activity. Therefore, the folding of multidomain proteins in the cell often proceeds with the assistance of various molecular chaperones. Because the intracellular environment is a crowded environment in which the concentration of macromolecules is estimated as 300–400 mg/mL, the risk of aggregation due to nonspecific molecular interactions exists. Folding of larger proteins and multidomain proteins often requires highly complex processes and, therefore, takes longer. Assembling multiple domains into one polypeptide chain is believed to be an evolutionary strategy to build up a protein with novel functions. However, a large proportion of the proteome consists of multidomain proteins: 40–65% of prokaryotic proteins and 65–80% of eukaryotic proteins. These studies have provided structural insights into folding intermediates and, thus, have expanded the understanding of the major folding pathways. Protein folding has been traditionally studied on small single-domain proteins, including BPTI, ribonuclease, and cytochrome c. To understand the mechanism of protein folding, experimental and computational studies have been performed over several decades. However, little is known about how proteins find efficient folding pathways and the process of structure formation. 
Despite the huge number of conformations possible, proteins often fold in seconds or even shorter periods (the Levinthal paradox). The newly synthesized protein searches for its own three-dimensional structure from myriad possibilities. However, the understanding of structure formation, folding, remains one of the biggest mysteries in protein science. Our results thus provide a global overview of the chaperone network properties in yeast, expanding our understanding of their functional diversity and their role in protein homeostasis.Significant advances in protein structure prediction have led to the easy availability of protein structural information. Further analysis with established cellular complexes revealed the involvement of R2TP chaperone in quaternary structure formation. Systematic analysis on the physicochemical properties for all chaperone interactors further revealed the presence of preferential domains and folds that are highly interactive with chaperones such as the WD40 repeat domain. Most notably, we observed Hsp10, Hsp70 Ssz1 chaperone, and Hsp90 cochaperone Cdc37 to be the main drivers of the network architecture. epistatic) interaction network obtained for chaperones was further analyzed, revealing that the global topological parameters of the resulting network have a more central role in mediating interactions in comparison to the rest of the proteins in the cell. Recently, we had carried out experiments that elucidated a comprehensive interaction network for the core 67 chaperones and 15 cochaperones in the budding yeast Saccharomyces cerevisiae. Molecular chaperones are typically promiscuous interacting proteins that function globally in the cell to maintain protein homeostasis.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |



 RSS Feed
RSS Feed
